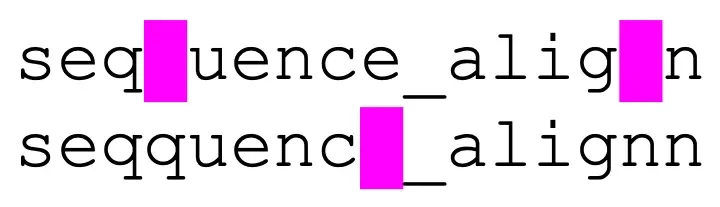Introducing sequence_align: An open-source Python + Rust toolkit for efficient sequence alignment
Kensho open-sources sequence_align, a Python + Rust toolkit for fast, scalable global sequence alignment on large vocabularies — without Python version constraints.
A universal step in evaluating the machine learning models that power Kensho Scribe is performing global sequence alignment. This step determines where differences exist between a reference sequence, such as words or speaker tags, and a sequence predicted by our model. However, given the scale of data that we work with at Kensho, this became a bottleneck due to time and memory complexity of global sequence alignment algorithms.
An example of an optimal alignment between the character sequences “sequence_align” and “seqquenc_alignn”.
After working with various other sequence alignment packages, we found that all of them had at least one of the following issues:
Too slow, requiring hours or even days to process evaluation sets even when parallelized
Too memory-intensive, restricting parallelization and/or requiring more expensive high-memory setups (e.g., AWS R5)
Too bioinformatics-specific, only working with single-character vocabularies like nucleotides instead of arbitrary string or integer values
Abandonware and/or only supporting older Python versions
Enter the sequence_align Python package. sequence_align is the only Python package on PyPi that addresses all of the above issues in our workflows, allowing us to quickly perform global sequence alignment at scale on sequences with arbitrarily large vocabularies without needing to downgrade to EOL Python versions.
Unlike several other toolkits available, we chose neither to write it in pure Python nor to use C extensions (e.g., Cython), two of the most common options for Python packages. Pure Python implementations are often much slower than those with bindings to a compiled language like C, but C extensions can often be quite verbose and have memory safety issues. Therefore, we decided to take a look at a newer option: PyO3, a cleverly named tool for creating Python bindings to Rust. Rust is a relatively new compiled language that offers similar speed benefits to C, but with excellent memory safety guarantees, so we decided to give it a shot!
Quick start
sequence_align is distributed via PyPi for Python 3.7+, making installation as simple as the following — no special setup required for cross-platform compatibility, Rust installation, etc.!
pip install sequence_alignPairwise sequence alignment algorithms are available in sequence_align.pairwise. Currently, two algorithms are implemented: the Needleman-Wunsch algorithm and Hirschberg’s algorithm. Needleman-Wunsch is commonly used for global sequence alignment, but suffers from the fact that it uses O(M*N) space, where M and N are the lengths of the two sequences being aligned. Hirschberg’s algorithm modifies Needleman-Wunsch to have the same time complexity (O(M*N)), but only use O(min{M, N}) space, making it an appealing option for memory-limited applications or extremely large sequences.
Using these algorithms is straightforward:
from sequence_align.pairwise import hirschberg, needleman_wunsch
# See https://en.wikipedia.org/wiki/Needleman%E2%80%93Wunsch_algorithm#/media/File:Needleman-Wunsch_pairwise_sequence_alignment.png
# Use Needleman-Wunsch default scores (match=1, mismatch=-1, indel=-1)
seq_a = ["G", "A", "T", "T", "A", "C", "A"]
seq_b = ["G", "C", "A", "T", "G", "C", "G"]
aligned_seq_a, aligned_seq_b = needleman_wunsch(
seq_a,
seq_b,
match_score=1.0,
mismatch_score=-1.0,
indel_score=-1.0,
gap="_",
)
# Expects ["G", "_", "A", "T", "T", "A", "C", "A"]
print(aligned_seq_a)
# Expects ["G", "C", "A", "_", "T", "G", "C", "G"]
print(aligned_seq_b)
# See https://en.wikipedia.org/wiki/Hirschberg%27s_algorithm#Example
seq_a = ["A", "G", "T", "A", "C", "G", "C", "A"]
seq_b = ["T", "A", "T", "G", "C"]
aligned_seq_a, aligned_seq_b = hirschberg(
seq_a,
seq_b,
match_score=2.0,
mismatch_score=-1.0,
indel_score=-2.0,
gap="_",
)
# Expects ["A", "G", "T", "A", "C", "G", "C", "A"]
print(aligned_seq_a)
# Expects ["_", "_", "T", "A", "T", "G", "C", "_"]
print(aligned_seq_b)Performance benchmarks
All tests below were conducted sequentially on an AWS R5.4 instance with 16 cores and 128 GB of memory. The pair of sequences for alignment consist of a character sequence of randomly selected A/C/G/T nucleotide bases along with another that is identical, except with 10% of the characters randomly perturbed by deletion, insertion of another randomly selected character after the entry, or replacement with a different randomly selected character.
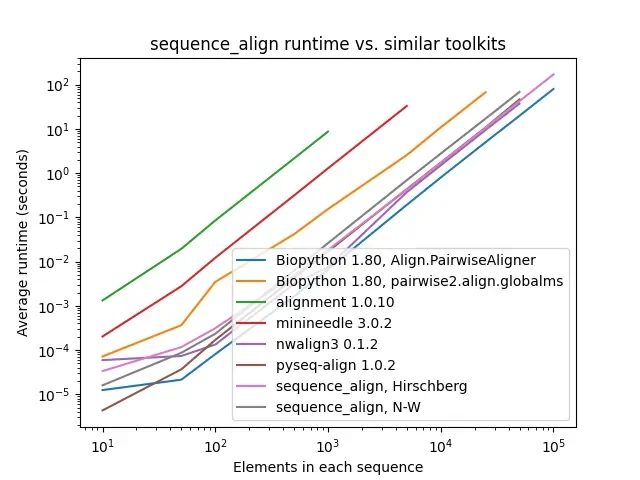
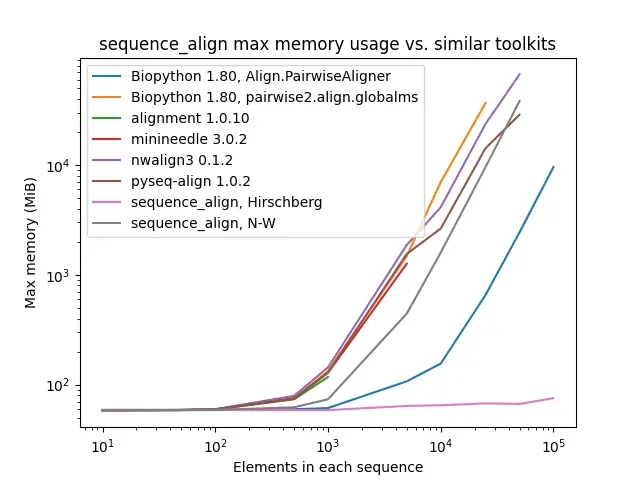
As one can see, while sequence_align is comparable to some other toolkits in terms of speed, its memory performance is best-in-class, even when compared to toolkits using the same algorithm, such as Needleman-Wunsch being used in pyseq-align.
Runtime comparison of toolkits tested. Some setups (e.g., minineedle) take prohibitively long at higher scales, so their lines terminate early.
Max memory usage comparison of toolkits tested. Some setups (e.g., Biopython pairwise2.align.globalms) run out of memory at higher scales, so their lines terminate early.
Concluding remarks
We hope that sequence_align will be a useful tool to the community across multiple fields, including natural language processing, bioinformatics and more. Please reach out to sequence-align-maintainer@kensho.com if you have any concerns or suggestions!
Kensho is an artificial intelligence company that builds solutions to uncover insights in messy and unstructured data. The services in Kensho’s AI toolkit, including Scribe, NERD, Extract and Link, enable critical workflows and empower businesses to make decisions with conviction.
Kensho encourages a growth mindset fueled by intellectual curiosity. We make it a priority to continually invest in our employees’ development and long-term success through benefits such as monthly knowledge days, a continual learning budget (which can be used to attend conferences like INTERSPEECH!), tuition reimbursement, and a wide range of development opportunities. Learn more about Kensho’s benefits and check out our open positions on our website at kensho.com.